Choosing a Cloning VectorVectors must be relatively small molecules for convenience of manipulation. Theymust be capable of prolific in a living cell, thereby enabling theof the inserted donor fragment.
Another important requirement isthat there be convenient restriction sites that can be used for insertion of theto be cloned. Unique sites are most useful because then the insert can betargeted to one site in the. It is also important that there be amechanism for easy identification and recovery of the molecule.There are numerous cloning vectors in current use, and the choice between themoften depends on the size of the DNA segment that needs to be cloned. We willconsider several commonly used types. PlasmidsAs described earlier, bacterial plasmids are small circular moleculesthat are distinct from, as well as additional to, the main bacterial. They replicate their DNA independently of the bacterialchromosome. Many different types of plasmids have been found in bacteria.The of any one within a species is generally sporadic;some cells have the plasmid, whereas others do not.
In Chapter 9, we encountered the Fplasmid, which confers certain types of conjugative behavior to cells ofE. The F plasmid can be used as a forcarrying large inserts, as we shall see in Chapter 12. However, theplasmids that are routinely used as vectors are those that carry genes fordrug resistance. The drug-resistance genes are useful because thedrug-resistant can be used to select not only for cellstransformed by plasmids, but also for vectors containing.Plasmids are also an efficient means of amplifying cloned DNA because thereare many copies per cell, as many as several hundred for some plasmids.Two vectors that have been extensively used in are shown inon the followingpage. These vectors are derived from natural plasmids, but both have beengenetically modified for convenient use as vectors.
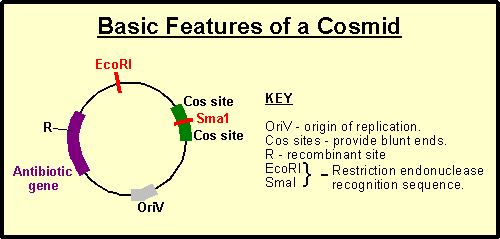
A cosmid is a type of hybrid plasmid that contains a Lambda phage cos sequence. Cosmids (cos sites + plasmid = cosmids) DNA sequences are originally from the lambda phage. They are often used as a cloning vector in genetic engineering. Cosmids can be used to build genomic libraries.
PlasmidpBR322 is simpler in structure; it has two drug-resistance genes,tet R andamp R. Both genes contain unique restrictiontarget sites that are useful in cloning. For example, could beinserted into the tet R.
Successfulinsertion will split and inactivate the tet Rgene, which then will no longer confer tetracycline resistance, and the cellwill be sensitive to that drug. Therefore, the cloning procedure would be tomix the samples of cut plasmid and donor DNA, transform bacteria, and selectfirst for ampicillin-resistant colonies, which must have been successfullytransformed by a plasmid molecule. Of the Amp R colonies, onlythose that prove to be tetracycline-sensitive have inserts; in other words,the Amp R Tet S colonies are the ones that containrecombinant DNA. Further experiments are needed to find the clones with thespecific insert required. Two plasmids designed as vectors for DNA cloning, showing generalstructure and restriction sites. Insertion into pBR322 isdetected by inactivation of one drug-resistance gene( tet R), indicated by thetet S (sensitive) phenotype.Insertion into pUC18The pUC is a more advanced, whose structure allows directvisual selection of colonies containing vectors with inserts.
Thekey element is a small part of the E. Coli β-galactosidase. Into this region has been inserted a piece of DNA called a, or multiplecloning site, which contains many unique restriction target sitesuseful for inserting donor fragments. The is in frametranslationally with the β-galactosidase fragment and does not interferewith its. The protocol uses recipient cells thatcontain a β-galactosidase gene lacking the fragment present on the plasmid.An unusual type of occurs in which the partial proteinscoded by the two fragments unite to form a functional β-galactosidase.colorless substrate for β-galactosidase called X-Gal is added to the,and the functional converts this substrate into a blue dye, whichcolors the blue. If donor DNA is inserted into the polylinker, theenzyme fragment borne on the vector is disrupted, no completeβ-galactosidase protein is formed, and the colony is white. Hence, selectionfor white Amp R colonies selects directly for vectors bearinginserts, and such colonies are isolated for further study.Plasmids that contain large inserts of tend to spontaneously losethe insert; therefore, plasmids are not useful for cloning DNA fragmentslarger than 20 kb.
Phage lambdaPhage λ is a convenient cloning for several reasons. First, λheads will selectively package a about 50 kb in length, and, aswill be seen, this property can be used to select for λ molecules withinserts of. The central part of the phage is not requiredfor or packaging of λ DNA molecules in E. Coli,so the central part can be cut out by using restriction enzymes anddiscarded.
The two “arms” are ligated to restriction-digested donor DNA. Thechimeric molecules can be either introduced into E. Colidirectly by or packaged into phage heads. In the invitro system, DNA and phage-head components are mixed together, andinfective λ phages form spontaneously. In either method,molecules with 10- to 15-kb inserts are the ones that will be mosteffectively packaged into phage heads, because this size of insertsubstitutes for the deleted central part of the phage genome and brings thetotal molecule size to 50 kb.
Therefore the presence of a phage onthe bacterial automatically signals the presence of recombinant phagebearing an insert.second useful property of a phage vector is that recombinant molecules areautomatically packaged into infective phage particles, which can beconveniently stored and handled experimentally. CosmidsCosmids are vectors that are hybrids of λ phages and plasmids, and theircan replicate in the cell like that of a or be packaged like that ofa.
However, cosmids can carry DNA inserts about three times as largeas those carried by λ itself (as large as about 45 kb). The key is that mostof the λ phage structure has been deleted, but the signal sequences thatpromote phage-head stuffing ( cos sites) remain. Thismodified structure enables phage heads to be stuffed with almost all. Cosmid DNA can be packaged into phage particles by using thesystem. Cloning by cosmids is illustrated in. Expression vectorsOne way of detecting a specific cloned is by detecting its proteinproduct expressed in the bacterial cell.
Therefore, in these cases, it isnecessary to be able to express the gene in bacteria; that is, to transcribeit and translate the mRNA into protein. Most cloning vectors do not permitexpression of cloned genes, but such expression is possible if specialvectors are used. However, because bacteria cannot process introns, thecloned sequences must be stripped of introns.
The cloned gene is insertednext to appropriate bacterial and start signals.Some expression vectors have been designed with restriction sites locatedjust next to a lac. These restrictionsites permit to be spliced into the for expression underthe control of the lac regulatory system. Making a DNA LibraryWe have learned that the most important goal of technology is toa particular or other genomic fragment of interest to the researcher.The approach used to clone a specific gene depends to a large degree on the genein question and on what is known about it. Generally, the procedures start witha sample of DNA such as eukaryotic genomic DNA. The next step is to obtain alarge collection of clones made from this original DNA sample. The collection ofclones is called a DNA. This step is sometimes referred toas “shotgun” cloning because the experimenter clones a large sample of fragmentsand hopes that one of the clones will contain a “hit”—the desired gene.
The taskthen is to find that particular clone.There are different types of libraries, categorized, first, according to whichis used and, second, according to the source of. Different cloningvectors carry different amounts of DNA, so the choice of vector forconstruction depends on the size of the (or other DNA sample) being madeinto the library.
Plasmid and vectors carry small amounts of DNA, so thesevectors are suitable for cloning genes from organisms with small genomes.Cosmids carry larger amounts of DNA, and other vectors such as YACs and BACs(see Chapter 12) carry the largestamounts of all. Ease of manipulation is another important factor in choosing avector.
Phage library is a suspension of phages. A or a libraryis a suspension of bacteria or a set of defined bacterial cultures stored intubes or microtiter dishes.The second important decision is whether to make a or a. CDNA, or, is syntheticmade from mRNA with the use of a special called originally isolated from retroviruses.
With the useof an mRNA as a, reverse transcriptase synthesizes a single-strandedDNA molecule that can then be used as a template for double-stranded DNAsynthesis. Because it ismade from mRNA, cDNA is devoid of both upstream and downstream regulatorysequences and of introns. This means that cDNA from eukaryotes can be translatedinto functional protein in bacteria—an important feature when expressingeukaryotic genes in bacterial hosts.
The synthesis of double-stranded cDNA from mRNA. A short oligo(dT)chain is hybridized to the poly(A) tail of an mRNA strand. Theoligo(dT) segment serves as a primer for the action of reverse transcriptase, which uses the mRNA as a template for the synthesisofThe choice between genomic and depends on the situation.
If a specificthat is active in a specific type of tissue in a plant or animal is beingsought, then it makes sense to use that tissue to prepare mRNA to be convertedinto cDNA and then make a from that sample. This library should beenriched for the gene in question. CDNA library is based on the regions of thetranscribed, so it will inevitably be smaller than a complete, which should contain all of the genome. Although genomic libraries arebigger, they do have the benefit of containing genes in their native form,including introns and regulatory sequences. If the purpose of constructing thelibrary is a prelude to cloning an entire genome, then a genomic library isnecessary at some stage.In some cases, it is possible to narrow down the genomic fraction used inconstruction, to more easily detect the desired. This approach is possibleif the experimenter already knows which contains the gene. Onetechnique used in mammalian is to sort the chromosomes withan instrument called a flow cytometer.
Suspension ofchromosomes is passed through the apparatus, which sorts the chromosomesaccording to size (this procedure is discussed in more detail in Chapter 12). The appropriatechromosomal fraction is then used to make the library.Another technique possible in organisms with small chromosomes is to fractionatewhole chromosomes by using pulsed field gel (PFGE).Electrophoresis is a general technique that fractionates nucleic acids orproteins according to size on gels under the influence of a strong electricfield. This type of procedure separates shorter fragments. PFGE is aspecialized type of electrophoresis useful for very long DNA molecules.
It usesseveral oscillating electric fields oriented in several different directions.This enables large DNA molecules such as whole chromosomes to snake through thegel to different positions according to their size. The appropriatecan be identified on the gel by probing with a chromosome-specific (seethe next subsection). Then the desired chromosome can be cut out, eluted fromthe gel, and used to make a chromosome-specific.How can an experimenter determine whether a is large enough to containany one unique sequence of interest with a reasonable degree of certainty? Thereare formulas for calculating the minimum number of clones needed, but a roughidea of the general order of magnitude of the library can be obtained simply bytaking the total size and dividing by the average size of the insertscarried by the being used. Generally, this number will be at leastdoubled, but it does provide a rough estimate of the magnitude of the job oflibrary construction. Identification of a specific in a is a two-step procedure.
First, coloniesor plaques of the library on a petri are transferred to an absorbentmembrane (often nitrocellulose) by simply laying the membrane on the surfaceof the. The membrane is peeled off, and colonies or plaques clingingto the surface are lysed and the denatured. The next step is tobathe the membrane with a solution of a that is specific for the DNAbeing sought.
The probe must be labeled either with radioactivity or afluorescent dye. Generally, the probe is itself a cloned piece of DNA thathas a sequence homologous to the desired. The probe DNA must bedenatured; it will then bind only to the DNA of the clone being sought. Theposition of a positive clone will become clear from the position of theconcentrated label, often as a spot on an.
Facing page: (a) A genomic library can be madeby cloning genes in λ bacteriophages. When a lawn of bacteria ona petri plate is infected by a large number of different hybridphages, each plaque in the lawn is inhabited by a single cloneof phagesWhere does the to make a come from?
The DNA can be from one ofseveral sources. One source is from tissue that expresses the ofinterest. The idea is that, because the mRNA of a gene is abundant, many ofthe cDNAs made from this tissue and inserted individually into vectors willvery likely be for the desired gene. For example, in mammalianreticulocytes, 90 percent of the mRNA is known to be transcribed from theβ-globin gene, so reticulocytes would be a good source of mRNA for making acDNA probe to find a genomic globin gene. In this case, awould be probed. The need for this kind of analysis depends on whichquestions are to be asked about the gene. If only the transcribed sequenceis of interest, then the cDNA itself could provide that informationjust as well.
However, if introns and control regions are needed, thegenomic clone must be obtained.Another source of for a might be a homologous from a relatedorganism. For example, if a certain gene has been cloned in the ascomycetefungus Neurospora, then it is very likely that this genecan be used as a probe to find the homologous gene in the related fungusPodospora. This method depends on the evolutionaryconservation of DNA sequences through time.
Even though the probe DNA andthe DNA of the desired might not be identical, they are often similarenough to promote hybridization. The method is jokingly called “clone byphone” because, if you can phone a colleague who has a clone of your gene ofinterest but from a related organism, then your job of cloning is maderelatively easy.Probe can be synthesized if the protein product of the of interestis known and an sequence has been obtained. Synthetic DNA probesare designed on the basis of knowledge of the, so an amino acidsequence merely has to be translated backward to obtain the DNA sequencethat encoded it. However, because of the redundancy of the code—in otherwords, the fact that most amino acids are coded by more than one—several possible DNA sequences could have encoded the protein inquestion. To get around this problem, a short stretch of amino acids withminimal redundancy is selected.
The sequence is calculated byusing the codon dictionary. The chemical DNA synthesizing reaction is astep-by-step process, so wherever in the sequence there are alternativenucleotides, a mixture of those alternative nucleotides is fed into thereaction and all possible DNA strands are synthesized. Shows an example in which there are fivepositions of redundancy, showing 2, 3, 2, 2, and 2 alternatives,respectively.
The reaction would make 2 × 3 × 2 × 2 × 2 = 48 strands at thesame time. This “cocktail” of oligonucleotides would be used as a.
Thecorrect strand within this cocktail would find the gene of interest. Twentynucleotides embody enough specificity to find one unique DNA sequence in the. Probes for finding proteinsIf the protein product of a is known and isolated in pure form, thenthis protein can be used to detect the of the corresponding gene in a. The process is described in.
An to the protein is prepared, and this antibodyis used to screen an. These libraries are made by usingexpression vectors designed to express high levels of a specific bacterialprotein. To make the library, is inserted into the in frame withthe bacterial protein, and the cells will make a fusion protein.
Membraneis laid over the surface of the and removed with an imprint ofcolonies. It is dried and bathed in a solution of the antibody. Positiveclones are revealed by making an antibody to the first antibody; the secondantibody is labeled by a radioactive or a chemical that willfluoresce or become a colored dye. By detecting the correct protein, theantibody effectively identifies the clone containing the gene that must havesynthesized that protein.
Finding the clone of interest by using antibody. An expression library made with phage derivative λgt11 is screened with aprotein-specific antibody. After the un-bound antibodies havebeen washed off the filter, the bound antibodies are visualizedthroughAt the beginning of the chapter, we asked how it might be possible to findthe for human albinism. It was in fact cloned by using an tothe that is known to be defective in this condition, the enzymetyrosinase. This enzyme, like any protein, can be purified by standardbiochemical procedures, and subsequently an antibody to the enzyme wasprepared in rabbits.
From tyrosinase-producing cells, mRNA was isolated andused to make. This cDNA was used to make an.The library was probed with the antibody to tyrosinase, and several positiveclones were detected. The cDNA in the positive clones was sequenced andfound to contain a gene whose exons total 1590 pairs. The cDNAwas used to a library of human genomic, and, in this process, theintact tyrosinase gene was found. It proved to have five exons and fourintrons.
This method depends on the ability to transform the donor organism, often a. We have already considered in prokaryotes (Chapters 2 and 9), but eukaryotes can betransformed, too. The procedure differs among eukaryotes, but generally somespecial treatment of recipient cells is required. For example, to transformfungi, generally the cell walls must be removed enzymatically. Let’s assume thatwe have isolated a that is relevant to some biological process thatinterests us. For the present purpose, we will assume that it is an auxotrophicin a fungus. We shall use from the to transform theauxotrophic mutant and then these recipient fungal cells on.
Fungal cells that contain the wild-type (from the wild-typeused to make the library) will transform the to prototrophyand allow growth on minimal medium.The reason that this method works is that the transformingfragment functionally complements the deficiency caused by the inthe recipient. It might seem at first that this view of is notthe same as the one developed in Chapter6; that is, the production of a wild-type from the union oftwo mutant genomes. However, the transforming contributes something thatthe recipient lacks (the wild-type allele being sought), and therecipient genome contributes something that the vector lacks (the entireremainder of the genome), so a type of complementation is involved.If the recipient is an organism in which vectors replicateautonomously (mainly bacteria and yeasts), then the transforming insert can berecovered simply by isolating the plasmid. However, as we shall see, in mosteukaryotic organisms the bacterial or cannot replicate and mustinsert into the to achieve stable transformation. In these cases, thetransforming fragment is relatively inaccessible and must be retrieved from thesuccessful in the. This method uses a library in which the clonesare laid out as a collection of numbered bacterial cultures in tubes ormicrotiter dishes.
Is isolated in bulk from all the strains in specificsubsets of the library, and transformation is attempted. By a process ofnarrowing down the library subsets that successfully transform, the clone withthe wild-type can be identified The process is illustrated in, using as an example theNeurospora trp3 discussed at the beginning of thechapter. In this case, a library was used. The cosmid must also carry agene that can be used to select for successful transformants of thefungus. Gene for hygromycin resistance is commonly used in fungi, which arenormally sensitive to this drug. Subsets of the cosmid library made fromwild-type DNA were used to transformtrp3 cells, and trp3 +clones were selected by plating transformed cells on containinghygromycin but lacking tryptophan. Colonies that grow are likely to contain thetrp3 + allele and are isolated from the.
Finding a cloned gene by using progressively smaller pooled DNAsamples in transformation. In this example, the quest is for thetrp3 gene of Neurospora (AfterJ. Fincham, Genetic Analysis. Copyright ©1995 by Blackwell.)In most cases, transformants are found to contain the carrying thewild-type inserted into one of the recipient’s chromosomes at a locationthat is different from the locus in the recipient.
This is calledectopic insertion. Less commonly, the transforming wild-type allele replaces theresident auxotrophic by a double-crossover-like process. The possible fates of transforming DNA. A donor wild-type alleleA + (cloned in a bacterialvector) transforms an A − recipient byone of three different types of insertion. Note: Therecipient is generally of the same species as the donor DNA,If a eukaryotic is cloned on a prokaryotic but a specific eukaryoticsequence is known that can act as an, this sequence can beadded to the vector. Then the vector will be able to replicate in both bacterialand eukaryotic cells, and insertion into the is not essential.
Thesetypes of vectors are called shuttle vectors because they can bemoved back and forth between different hosts. Without an origin of replication,the must integrate into the eukaryotic chromosome toeffect stable. Positional CloningInformation about a ’s position in the can be used to circumvent thehard work of assaying an entire to find the of interest.Positional cloning is a term that can be applied to any methodthat makes use of such information. Often both probing and arepart of positional cloning.
Common starting point is the availability ofanother cloned gene or other known to be closely linked to the gene beingsought. The linked marker acts as the departure point in a process, called, that willterminate at the target gene. Summarizes the procedure of. End fragmentsof a clone of the linked marker are used as probes to select other clones fromthe library. These probes will detect clones of regions that overlap withthe initial clone. Restriction maps (pages 327–329) are made of the DNA of thissecond set of clones, and, again, outward fragments are used for a new round ofselection of overlapping clones from the library.
Hence the walking processmoves outward in two directions from the start site. Each clone can be sequencedor otherwise tested, depending on the intent of the exploration. Chromosome walking. One recombinant phage obtained from a phagelibrary made by the partial EcoRI digest of aeukaryotic genome can be used to isolate another recombinant phagecontaining a neighboring segment of eukaryotic DNA, as described inthe text.Sometimes a large insert that is known to contain the linked will alsoluckily contain the sought, and subcloning and will narrowdown the appropriate region of the. The availability of a large number ofneutral markers (restriction fragment length polymorphisms) dispersedthroughout most genomes has provided many useful start points. Positionalcloning has been particularly useful for cloning human genes, many of which haveno known biochemical function and cannot be easily selected. The human gene for, mentioned at the beginningof the chapter, was cloned by, and we shall examine itscloning in more detail in Chapter12.

For any case of chromosome walking, there must be some type ofcriterion to assess each step of the walk for the gene of interest, and thesecriteria depend on the individual gene concerned. Cloning a Gene by TaggingTagging is a cloning method that zeros in on the desired directly byinducing a in that gene by using a specific piece of as aninsertional. The specific sequence is then used as a tag to recover thegene. The approach is summarized in. One type of tag is transforming DNA. When exogenous DNA isadded by or by other methods such as injection, it can integrateinto the and become part of the. Ectopic integration is randomthroughout the genome, and apparently no segment of chromosomal DNA is immune tointegration.
When integration takes place within or near a gene, the integratingfragment acts as a mutagen, disrupting the function of the gene. This propertycan be used to good advantage. Suppose that we use a specific cloned genex + and transform x − cells of thedonor organism into x +. Many of the x + transformants willbe for the genes into which the transforming DNA has insertedectopically. Subset of such x + cells will be mutant for the targetgene a +, the gene of interest, and will be ofa −. Hence among the x + transformants,a − phenotypes are identified.
The next step is to thetransformants to determine if the a − phenotypesegregates with x +. If it does, the mutation is likely to have beencaused by the integration of the fragment containingx +. The DNA of this mutant is used toconstruct a, and gene x + can be used as ato recover the of the disrupted a gene.
To recoverthe intact wild-type a gene, a fragment of the disrupteda gene sequence is used in another round of probing, thistime with a wild-type library. Using DNA insertion as a tag for marking and recovering a gene fromthe genome. The tag DNA can be transforming DNA or an endogenoustransposon (movable element).similar approach uses transposons as tags. Transposons are naturally mobilefragments found in many organisms. When they move, they can insert anywhere inthe. If they insert into or near a, they can create a.(Transposons are described in more detail in Chapter 13). In a containing an active,mutants for the desired gene are selected.
Many of these mutants will be causedby the insertion of the transposon. This line is used to make a.A cloned part of the transposon DNA can then be used as a tag to recover thegene, in a manner similar to that shown in.
